How transcript decoding in imaging based spatial transcriptomics like 10x Xenium work
Published:
Summarizing my understanding of the transcript decoding step in 10x xenium, which also applies for the most part to other imaging based spatial transcriptomics technologies like Vizgen MERSCOPE and NanoString CosMx.
The primary purpose of this blog is to just summarise my understanding so that this can be useful for my reference later on. I don’t guarantee the maintenance of accuracy of this information, especially as time passes and newer versions of these technologies come.
Acknowledgment: This content is heavily inspired by and draws upon the excellent resource from 10x Genomics: Xenium GEX Workshop Guide. All original material belongs to 10x Genomics, and this blog is intended as a derivative educational resource that complements their work. No copyright infringement is intended.
Introduction
- Its fascinating how using simple imaging of tissues, technologies are able to capture hundreds of genes in the samples, especially because one image is restricted by a very small number of channels.
- Multiple cycles of imaging with fluorescent probe binding and washing allow to capture a series of sequences of colors which let us decode what transcript was there at each point in the tissue. However, one thing to note here is that with more cycles, multiple issues arise (like more uncertainty in transcript decoding, and more noise in imaging), which cumulatively limit the number of imaging cycles we can have in the tissue.
- Here we talk about how we are able to capture hundreds of transcripts in image based spatial transcriptomics technologies.
Basic background of the chemistry upstream of transcript decoding
- This is the part where different imaging based spatial transcriptomics technologies differ.
- Sample is taken, processed to be able to accept the RNA binding probes: meaning mounted on slide, fixed and permeabilized.
- Padlock probes are allowed to bind to the sample.
- Washing is done to remove unbound probes.
- Rolling circle amplification is done.
- Cycles of fluorescent probe binding, imaging and washing are done.
Transcript Decoding
Refers to the process of inferring what transcript was present at a location, given the sequence of colors (or empty fluorescence) across the multiple cycles of imaging.
Codebook and Codewords
- Associated to each gene (or more explicitly, probe), we have a sequence of colors that are expected to be observed at the location of a transcript of that gene through the cycles. This is called the codeword for that gene. eg:
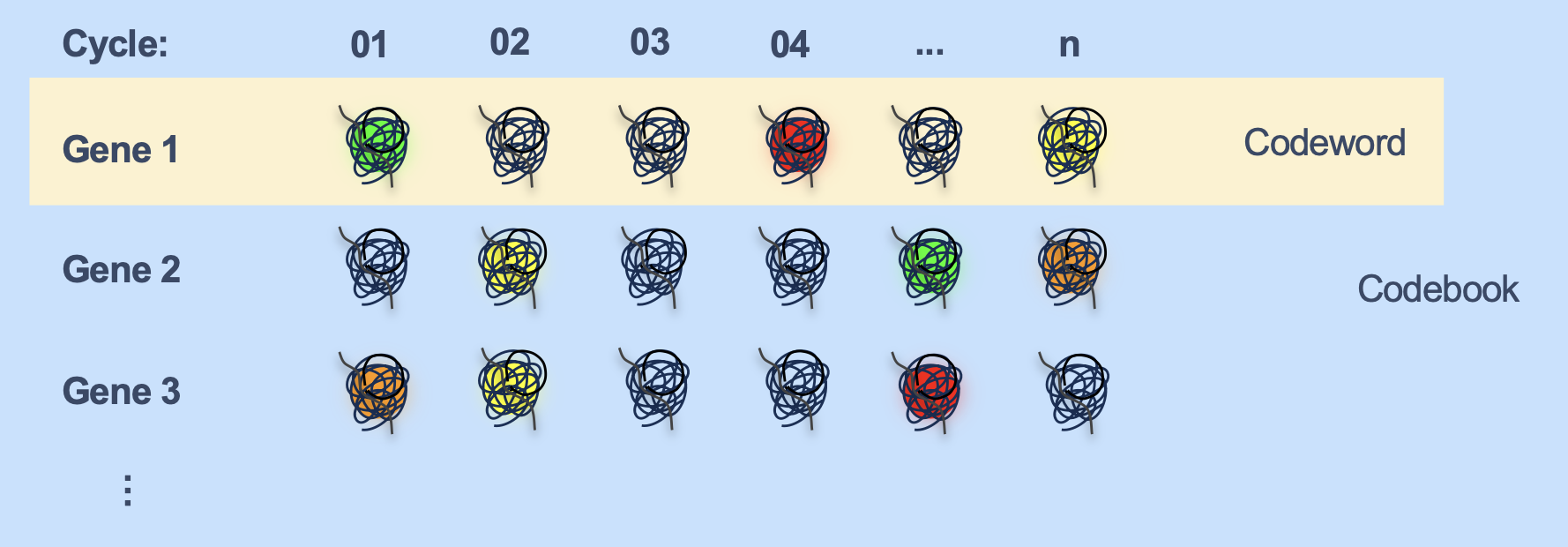
- The collection of all genes’ codewords is called the codebook.
Sparse encoding
If we have 3 different genes, whose expected color in a particular cycle is the same (say green), we may get something like this:
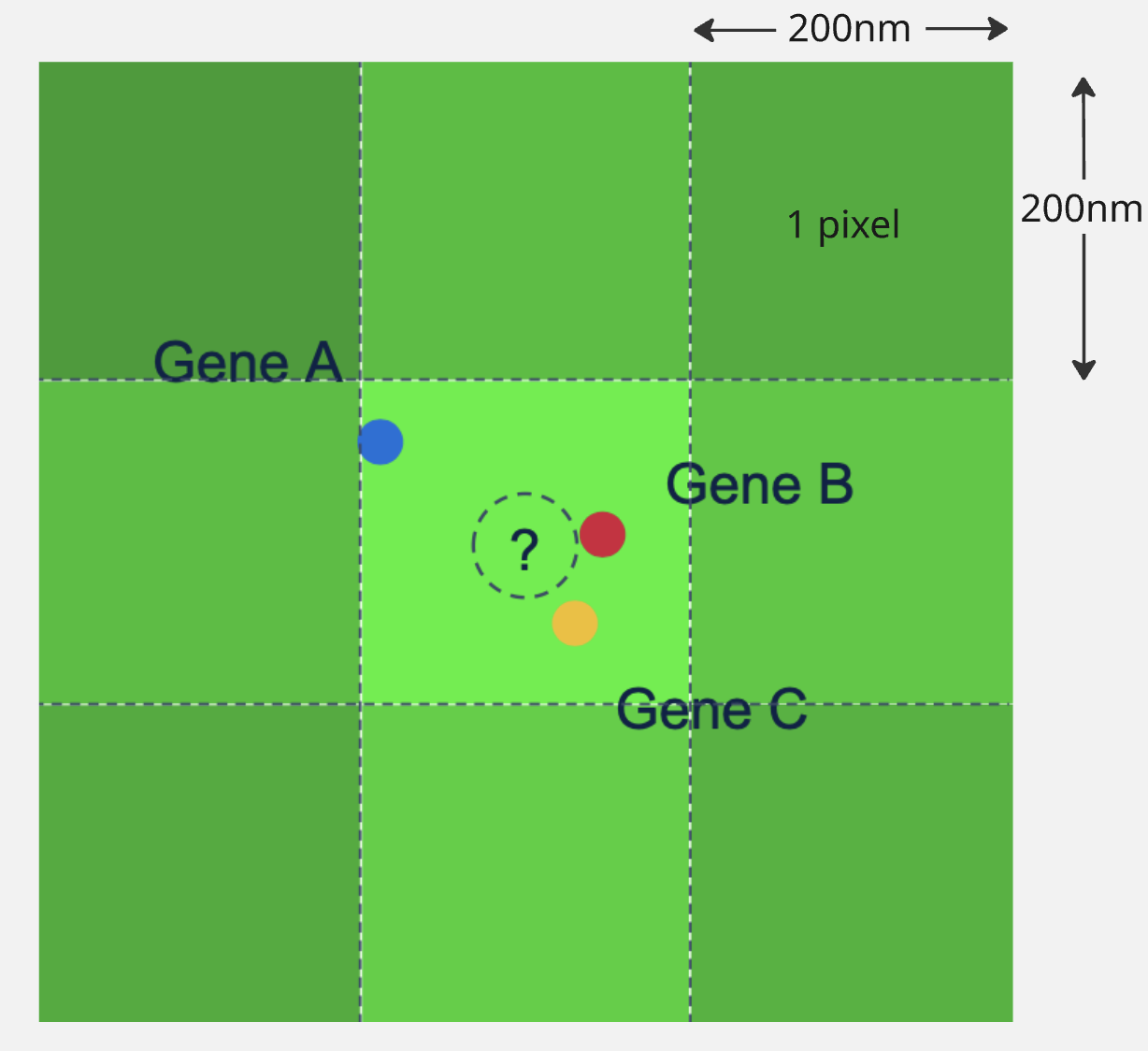
- This will appear as a single green spot in the image, making it difficult to determine that there were 3 transcripts there, all emitting the same green color.
- Sparse encoding helps to tackle this. Sparse encoding refers to the fact that a given gene’s codeword doesn’t contain more than a certain number of color signals, which is usually 5. The signal in the remaining positions in the codewords are empty signals.
- Since that leads to more empty signal positions in each codeword, it becomes easier to decipher the number of transcripts that would have given rise to the observed pixel intensity in the image.
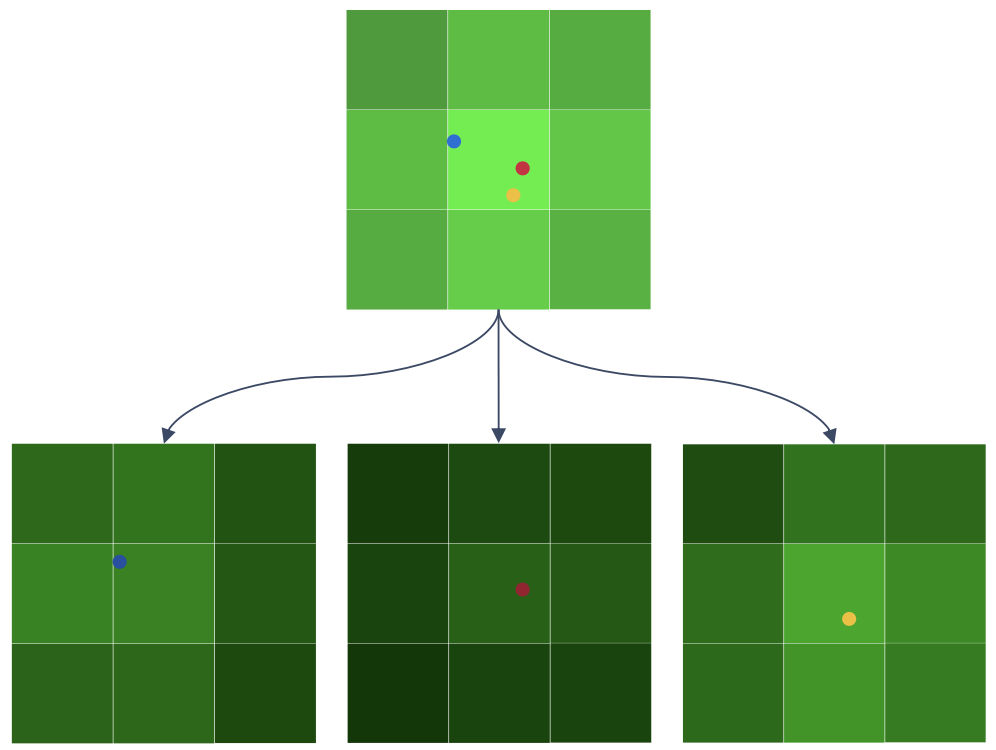
- Since the v1 chemistry of xenium (~300 genes) has 15 imaging cycles, ~10 positions in the codeword of each gene are empty.
- For example, gene VIM has the codeword: AEAEEDECEEEBEEE, where E is empty. Its detection at a spot in the tissue would look like:
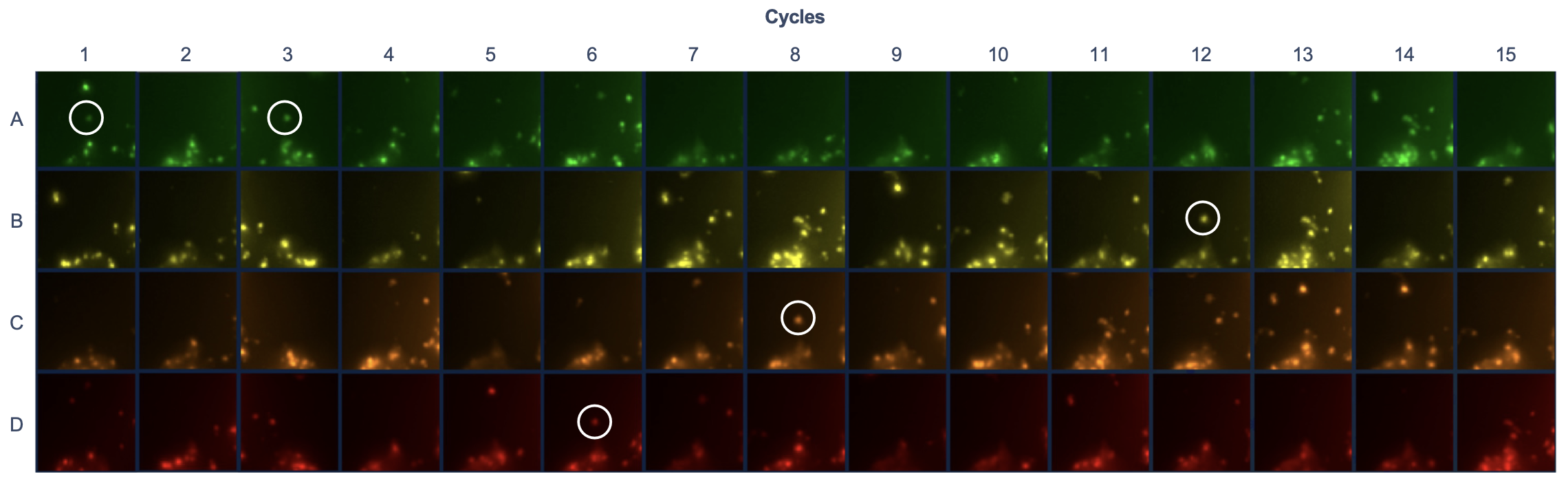
How the fluorescent probes (and padlock probes’ middle sequence) would have been, to be able to get the codewords for all genes in imaging
- Here is my understanding of how the fluorescent and padlock probes’ nucleotide sequence would be roughly designed so that we get the appropriate codeword for each gene.
- Likely, each color channel’s fluorescent probes for each imaging cycle has a unique nucleotide sequence, through which it binds to the padlock probe, because if not, then we would be wasting the information that could have been obtained through one channel of one imaging cycle.
- The barcodes of the fluorescent probes through which they bind to the padlock probes would remain constant.
- The middle part of a particular padlock probe would be designed such that it is able to bind to all the fluorescent probes’ barcode sequence to which that padlock probe is supposed to bind (the specific color channels and imaging cycles).
- If the length of the barcode of a fluorescent probe that binds to the padlock probe is n, then the total nucleotide sequence of padlock probes meant for binding fluorescent probes would be around 5n (assuming each padlock probe’s codeword has 5 colors and remaining empty signals).
Below is an example of a case for how the sequence for the middle part of the padlock probe would look like and the (fixed) sequence of the fluorescent probes would look like, if n were 4.
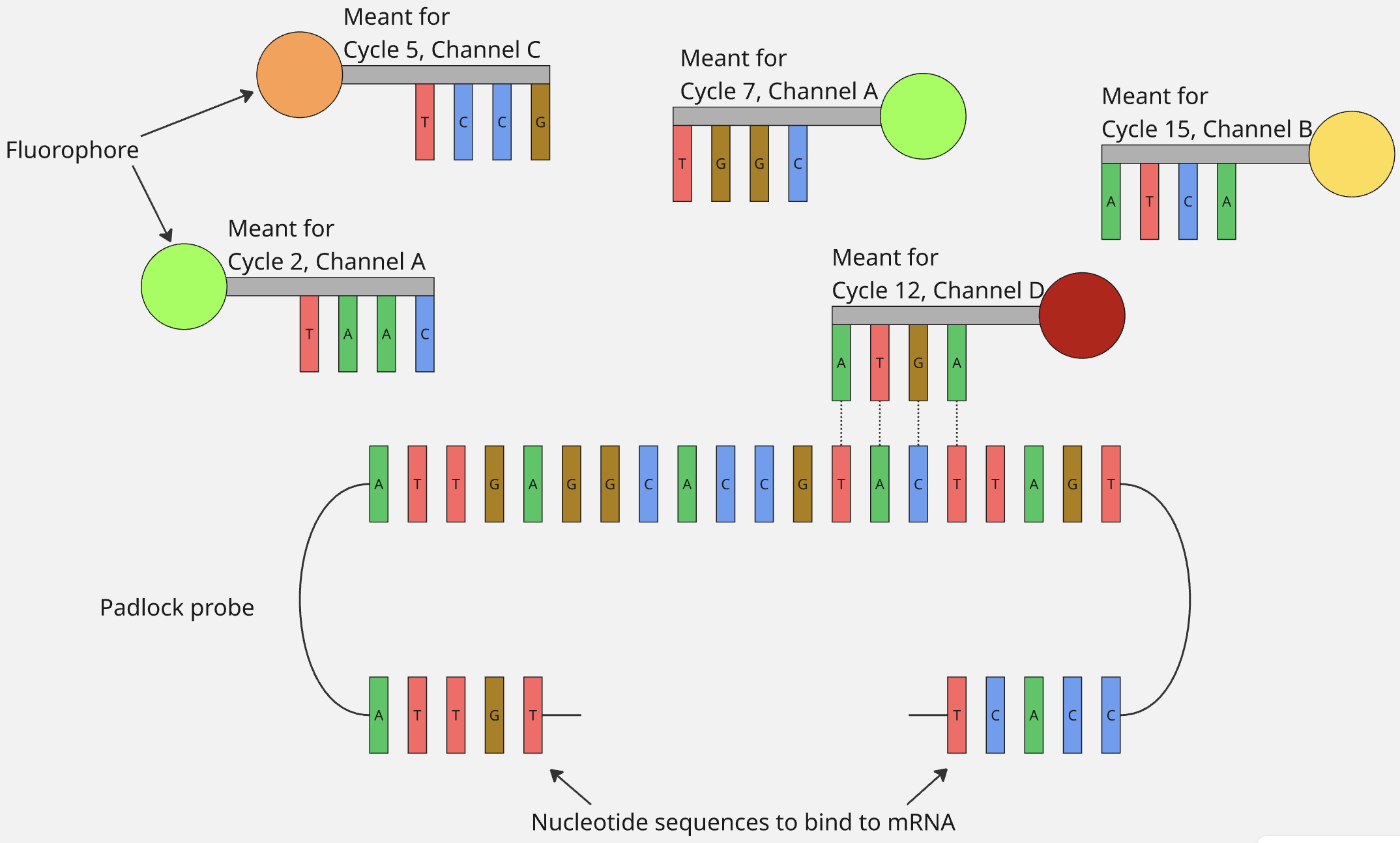
- When panels with custom genes are designed, then according to what codeword that gene is supposed to have, the nucleotide sequence of the middle part of its padlock probe is designed so that appropriate channel’s fluorescent probes bind to it during the appropriate imaging cycles.
- Again, this is my guess about this part, and I am not a 100% sure about this.
Quality score
- To assign a transcript at a location, Xenium decoding uses a probabilistic model that takes signal intensities, similarity to known codewords, and other attributes into account.
Using the probability of error obtained that way, a Q-score is computed as such: \(Q_{score}=-10\log_{10}\left(P_{err}\right)\).
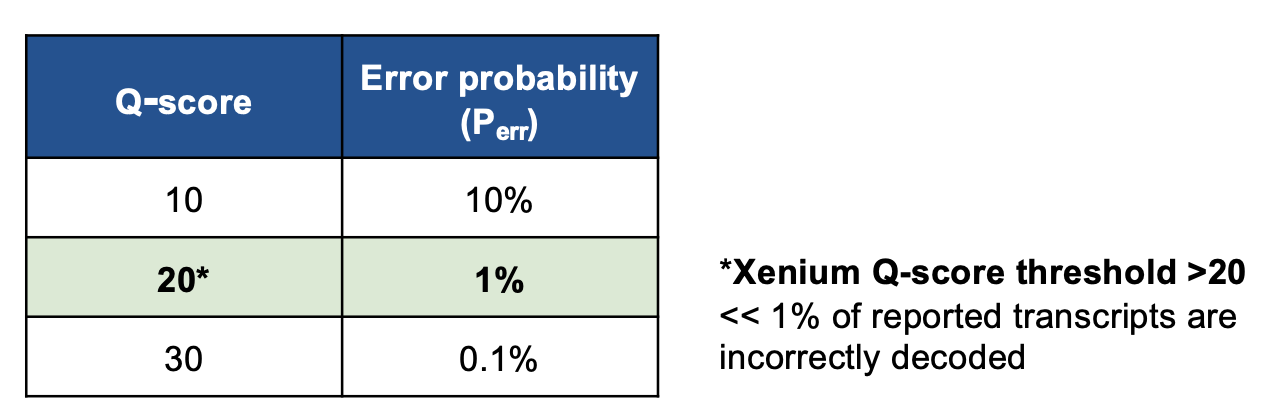

Negative Controls
- Negative Control Probes: Probes corresponding to transcripts which shouldn’t be present in our tissue. If they are detected highly, it should be interpreted as: “something that shouldn’t have bound, appears to have bounded to the tissue”. This indicates nonspecific binding or potential assay issues, meaning a significant number of detected transcripts are likely false positives rather than true biological signal. Common causes are: poor sample quality (eg. degraded RNA), incorrect experimental conditions (eg. low temperature washes), or specific tissue types like fresh-frozen.
- Negative Control Codewords: These are codewords corresponding to which there is no physical padlock probe in the experiment. If they are detected highly, it should be interpreted as: “Codeword corresponding to a padlock probe that wasn’t present, is detected”. This indicates possible issues with imaging, transcript decoding or autofluorescence.
- Unassigned Codewords: These are the codewords which aren’t assigned to any probe in the panel. Their purpose is to indicate the unused capacity from the panel. They are different from Negative Control Codewords in just that Negative Control Codewords are specifically chosen such that they can act as useful controls for the experiment (using parameters like similarity to existing gene codewords etc.), whereas unassigned codewords are simply unused codewords.
